


















When calling variants with the Basic Variant Detection, Low Frequency Variant Detection, or Fixed Ploidy Variant Detection tool and using the “Restrict calling to target regions” option, the following problems exist:
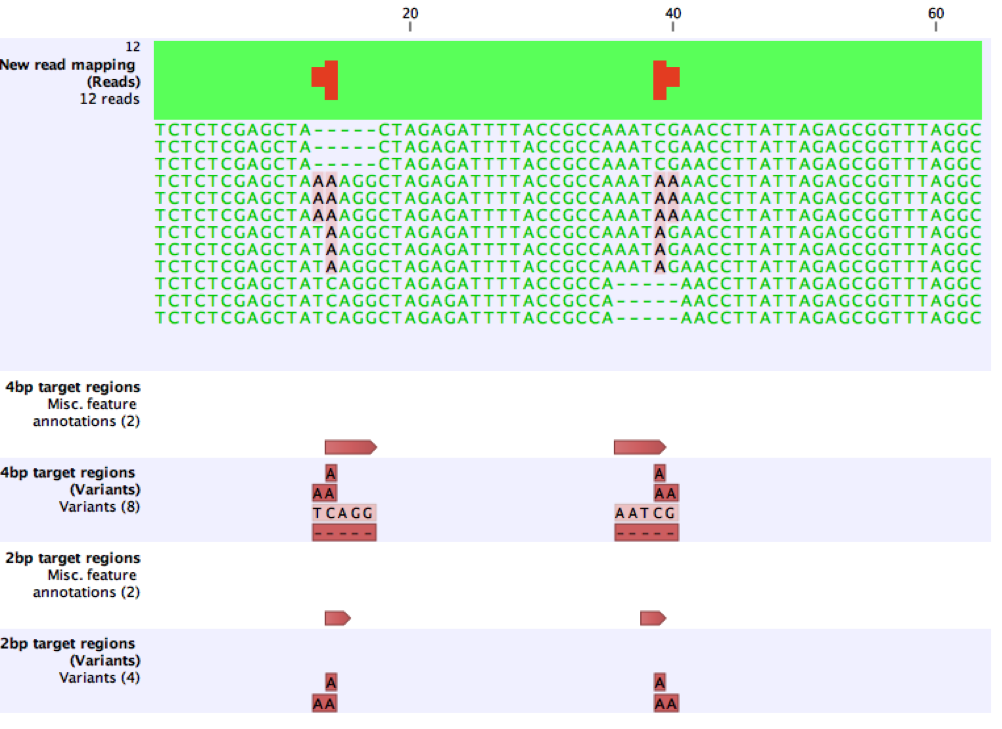
The effect of these issues are illustrated using a small, synthetic example. Here, 4bp target regions are provided, and six variants are called as expected. After reducing the target regions to 2bp, the 2bp MNVs, AG (left) and CA (right) are not called because they exactly match the coordinates of the target regions. When the target regions are further reduced to 1bp no variants are called.
This issue was fixed in CLC Genomics Workbench 12.0.2 and CLC Genomics Server 11.0.2.