


















Are you getting the most our of your RNA-Seq data from plant, animal, or microbial model organisms? Or are you investigating the transcriptome of an organism without reference genome?
Throughout this Spring we will be offering you information around RNA-Seq. Check out our scientific spotlight, featuring scientific applications of RNA-Seq including a recent article published by our customer Dr. Yasuda.
Dr. Masayuki Yasuda and colleagues at Department of Ophthalmology, Tohoku University Graduate School of Medicine, Sendai, Japan have investigated molecular mechanisms involved in RGC death by use of RNA-Sequencing technology and during their studies, QIAGEN products were used from sample to insight.
Glaucoma is a leading cause of blindness worldwide and thus an important disease to understand. The disease is characterized by retinal ganglion cell (RGC) death caused by axonal injury and in glaucoma patients the number of RGCs decreases due to axonal degeneration, resulting in visual dysfunction.
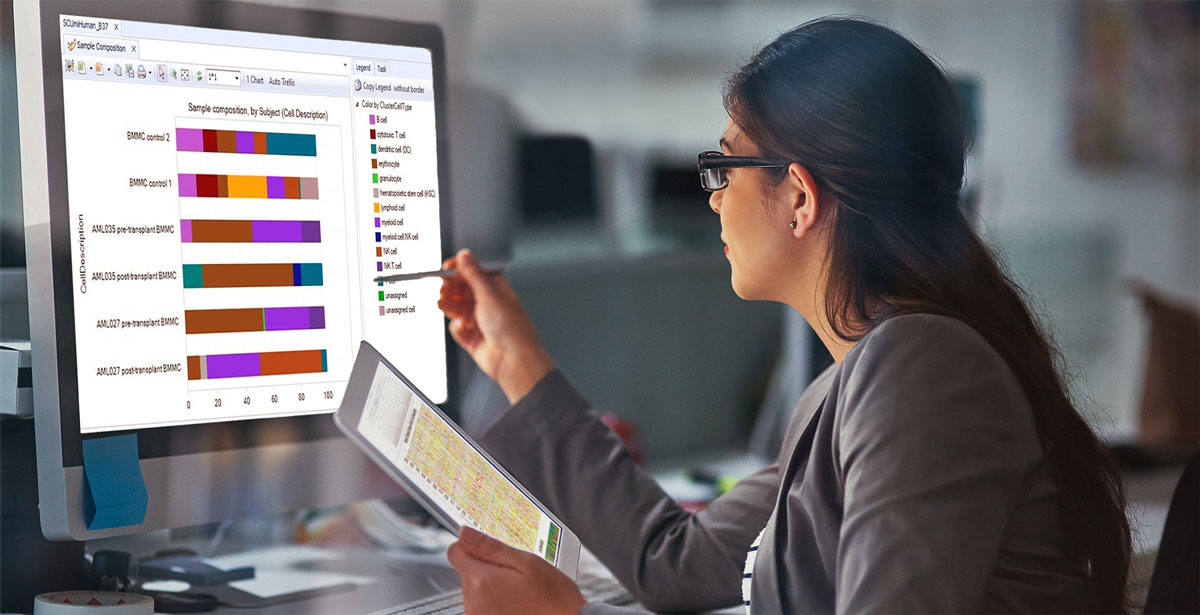
RNA-Seq was used to investigate the entire retinal transcriptome profile in the early stages of post-axonal injury. Sequencing data was imported to CLC Genomics Workbench for RNA-Seq, gene expression and statistical analysis.
Dr. Masayuki Yasuda says about their use of CLC Genomics Workbench:
“No one in my lab has ever experienced such high-throughput sequence data analysis. RNA-seq in my paper was my first high-throughput sequencing data analysis experience. Data analysis was not at all confusing and I could obtain very good results within a short period. The user interface is simple and customer support was very quick, that was very helpful. Also the detailed user manual was very informative. It would be great if I can do splice isoform analysis and pathway analysis in GWB as well.”
A pathway analysis revealed that the most significant biofunction in axonal injury was the “Cell Death and Survival” pathway. Pathway and global functional analyses were performed using IPA software predicting a key upstream transcription factor and a significant upstream regulator.
“Creating neuro protective drugs against optic nerve disorders is my research purpose. The challenge after extracting the list of differential expression genes from CLC Genomics Workbench was interpretation. How to find my next target for further analysis was a big challenge and I had thought a lot about it. At that time I remember that my colleague showed how to examine the mechanism of neuron apoptosis using pathway analysis. Coincidentally there was a seminar at core lab facility showing IPA, so I thought I could use this. IPA was a very powerful tool to find which pathway I should examine further. Especially upstream regulator analysis was very useful", says Dr. Masayuki Yasuda.
The transcriptomic approach relying on RNA-Seq, seems to be powerful and effective. This study may not only provide new insights into the molecular mechanisms underlying axonal damage but may also help in research aimed at the discovery of new biomarkers and therapeutic targets for a variety of ocular diseases.