


















Whole-exome sequencing (WES) using next-generation sequencing (NGS) technology is a powerful tool for investigating variants linked to genetic disease. It provides a high-resolution, unbiased view across the entire exome to discover causative variants of inherited disorders. However, the vast amounts of data produced by WES require comprehensive data analysis tools that can efficiently translate the raw sequencing data into meaningful, interpretable results. To address these challenges, QIAGEN Digital Insights offers QCI Interpret Translational.
QCI Interpret Translational is a NGS variant assessment software solution that enables rapid, evidence-powered variant annotation, filtering, and triage for human exome, genome, and large cohort sequencing data. QCI Interpret Translational rapidly identifies the most compelling disease variants in human sequencing data by combining powerful analytical tools and unparalleled content from the QIAGEN Knowledge Base. The QIAGEN Knowledge Base is the industry’s largest collection of biological and clinical findings, with roughly 2,000,000 unique variants expertly curated from over 300,000 scientific articles, including 140,000 variants connected to the top 200 newborn/carrier screening genes.
QCI Interpret Translational enables:
QCI Interpret Translational compiles all gene variants within a dataset and enables this list to be quickly narrowed down through an interactive series of filters. This Interactive Filter Cascade can be adapted to reflect selection criteria of interest and their importance to the research question at hand (Figure 1).
Figure 1. Rapid prioritization of variants. A list of gene variants relevant to an analyzed dataset can be drilled down to those most relevant to the research question by defining a series of filters that reflect the most important selection criteria.
Filters of the QCI Interpret Translational are based on information in the QIAGEN Knowledge Base, including manually curated primary literature on human mutations in patients with particular diseases or abnormal phenotypes. These filters offer a broad portfolio of selection criteria that go above and beyond identifying variants that impact symptoms, pathways, processes and genes implicated in drug response or disease progression.
Additional biologically relevant filters allow, for example, identification of diseases consistent with clinical features and deleterious variants in medical genomes, or use of single and bidirectional statistical burden tests to find genes or pathways with significantly more deleterious variants in one study group compared to another.
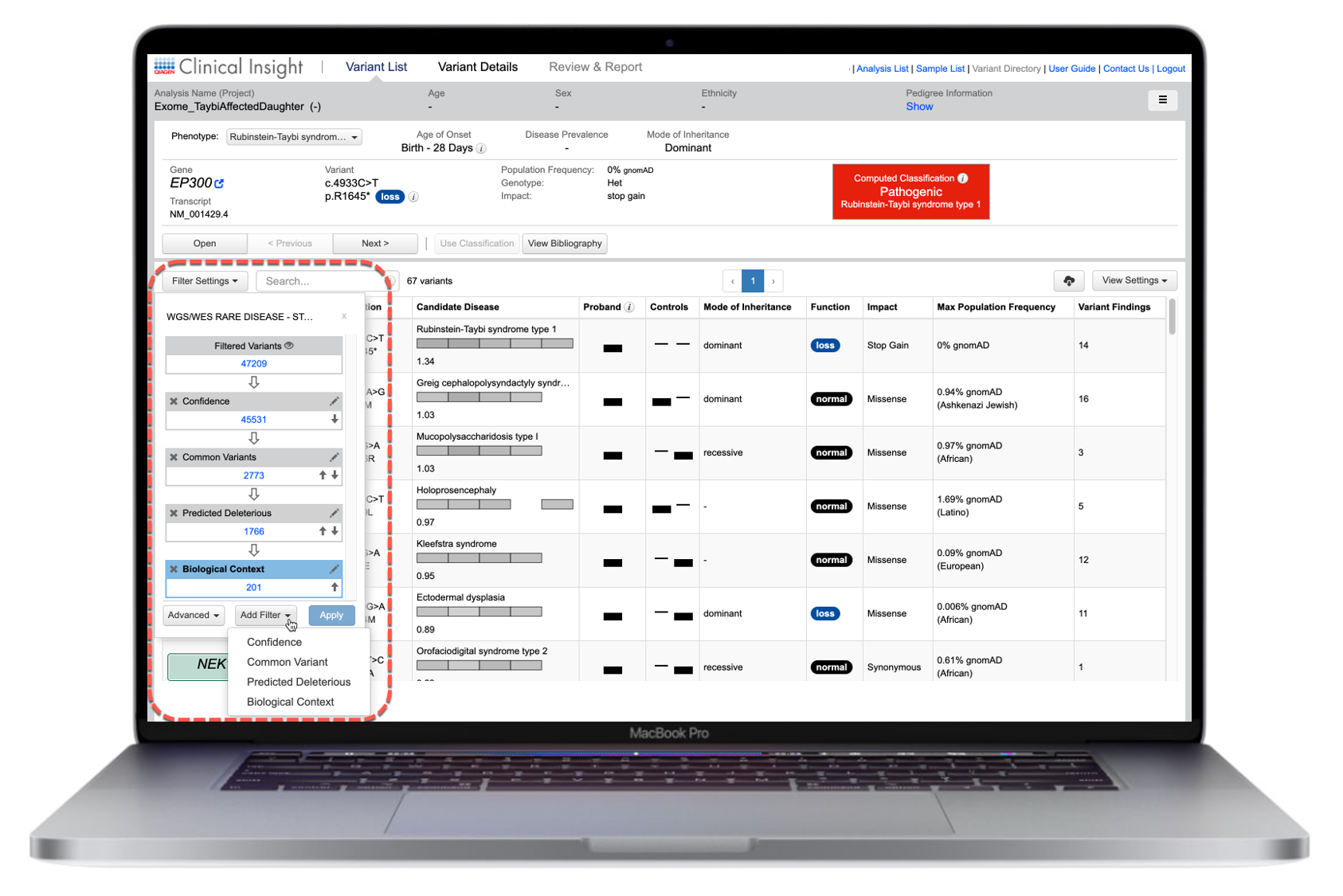
Users can access 5 different filters to triage variants with granular control (Figure 2):
Figure 2. Customizable dynamic variant filter. A list of gene variants relevant to an analyzed dataset can be drilled down to those most relevant to the research question by defining a series of filters that reflect the most important selection criteria.
Watch a video tutorial to learn how to use the Interactive Filter Cascade in QCI Interpret Translational here.