


















QCI Interpret is panel- and sequencer-agnostic and can be used with any secondary analysis platform.
The QCI Interpret workflow starts with a variant calling format (VCF) file and it is compatible with any NGS platform. You can chose to upload multiple-sample VCF files for different patients, or multiple single sample VCF files for samples from the same patient. Manually enter metadata to contextualize the uploaded VCF file in the hereditary workflow.
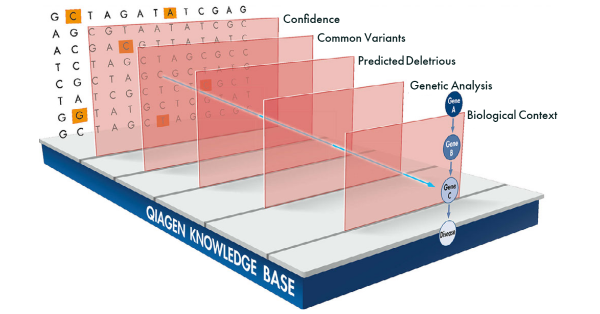 Use QCI Interpret's proprietary Interactive Filter Cascade (view filter here) to access 5 different filters to triage variants with granular control:
Use QCI Interpret's proprietary Interactive Filter Cascade (view filter here) to access 5 different filters to triage variants with granular control:
Pathogenicity classifications in QCI Interpret are accompanied by clear visibility into the evidence and criteria supporting the classifications. In addition, you can manually add a rationale for each of the rules triggered if desired and adjust the strength of the final assessment if additional data is available (see how here).
Review the literature as an essential step in the variant assessment process
QCI Interpret provides the most up-to-date, expertly curated extensive bibliography coverage (published clinical studies, including treatment and prognostic studies, functional studies, review articles, and other types) with multiple lines of evidence linking variants to a disease. Bibliography is categorized by article type for every variant detected and includes all findings that have been curated from the published literature as well as findings that have been imported from specific databases (view sample bibliography here).
Generate a final clinical report that is patient-specific and includes clinically relevant variants, interpretations, and references specified throughout the assessment process.

View full sample report for Hereditary Disorders here