
QCI Interpret for Hereditary Disease and Family Planning
Genetic testing and expanded carrier screening for rare and inherited diseases pose a unique challenge: accurately interpreting novel variants with limited supporting evidence. That’s why we developed QIAGEN Clinical Insight (QCI™) Interpret. Software designed to classify and interpret pathogenic variants in hereditary diseases, QCI Interpret taps into the power of the QIAGEN Knowledge Base, the industry’s largest, most up-to-date clinical database with information on over 750,000 human samples. With QCI Interpret, you can minimize interpretation risks and maximize your lab’s performance.
Home > QCI for Hereditary Disease
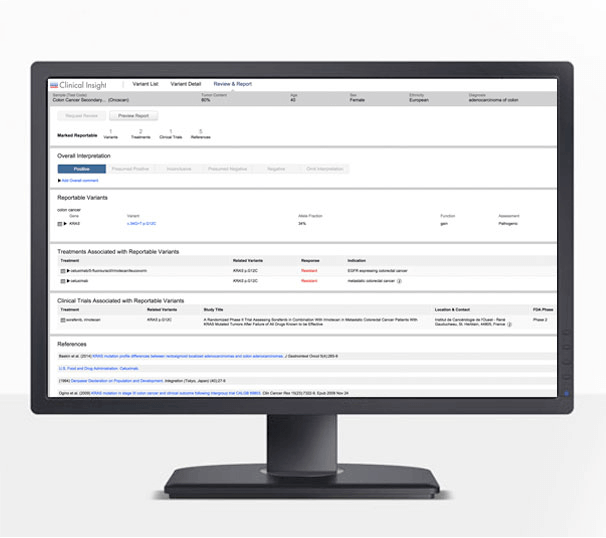
Growth powered by scalable insights
QCI Interpret replaces your complex and manual clinical NGS variant research with a scalable interpretation workflow alternative that helps grow your indication menu and volume. With QCI Interpret you can:
- Increase operational efficiency associated with scoring NGS variants by reducing time, costs, and complexity to classify variants.
- Focus interpretation time on high-risk variants that better inform genetic-profile specific patient care plans.
- Build your own experience-based database with variants assessed, increasing speed and accuracy with subsequent interpretations.
Reliable curation with superior depth
QCI Interpret is an efficiency tool that helps you identify pathogenic variants fast with results you can trust, all in one secure web application. With QCI Interpret you get:
Computed classifications that are based on rules published in ACMG professional medical guidelines for sequence variant interpretation.
Manually curated clinical case counts with clickable hyperlinks direct to source materials for easy retrieval and review.
Simple, clear, and easy-to-use configurable reports that also include bibliographic reference citations.

Request a QCI demo
Find out if QCI interpret for Hereditary Disease fits your NGS interpretation and reporting needs. Speak to a representative from QIAGEN Bioinformatics to take a closer look at a few representative data sets of variants that have been through a QCI Interpret workflow. Explore the easy-to-use interface, familiarize yourself with the various powerful features, see for yourself the detailed bibliography functionality, and practice building a comprehensive report.
Request your free, no obligation, demonstration of QCI™ Interpret for Hereditary Disease today.
*QCI Interpret is an evidence-based decision support software intended as an aid in the interpretation of variants observed in genomic sequencing data. The software evaluates genomic variants in the context of published biomedical literature, professional association guidelines, publicly available databases and annotations, drug labels, and clinical-trials. Based on this evaluation, the software proposes a classification to aid in the interpretation of observed variants. The software is NOT intended as a primary diagnostic tool by physicians or to be used as a substitute for professional healthcare advice. Each laboratory is responsible for ensuring compliance with applicable international, national, and local clinical laboratory regulations and other specific accreditations requirements.